Research interests of the Jack Vanden Heuvel Laboratory at Penn State.
Members of the NR superfamily act as intracellular transcription factors that directly regulate gene expression in response to lipophilic molecules. They affect a wide variety of functions, including fatty acid metabolism, reproductive development, and detoxification of foreign substances. To date, over 300 NRs have been cloned, many with unknown endogenous ligands (orphan receptors). Phylogenic analysis has shown six subfamilies (NR1-6) with various groups and individual genes. Several NRs have evolved to respond to dietary lipids and include the fatty acid receptors PPAR, LXR, RXR and FXR.
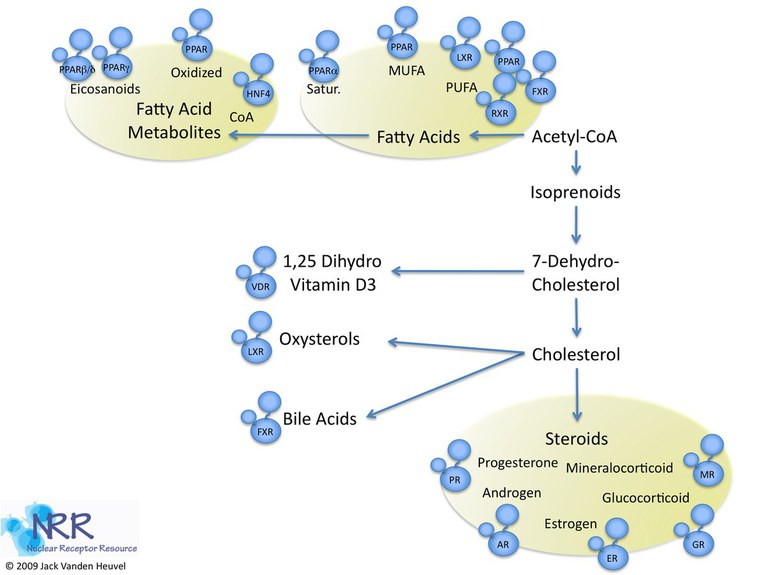
Figure 1
The receptors shown in Figure 1 may be considered constituents of a large group of NRs, the "metabolic nuclear receptors" which act as overall sensors of metabolic intermediates, xenobiotics and compounds in the diet and allow cells to respond to environmental changes by inducing the appropriate metabolic genes and pathways. Most NRs regulate gene expression in predominantly the same fashion (Figure 2).
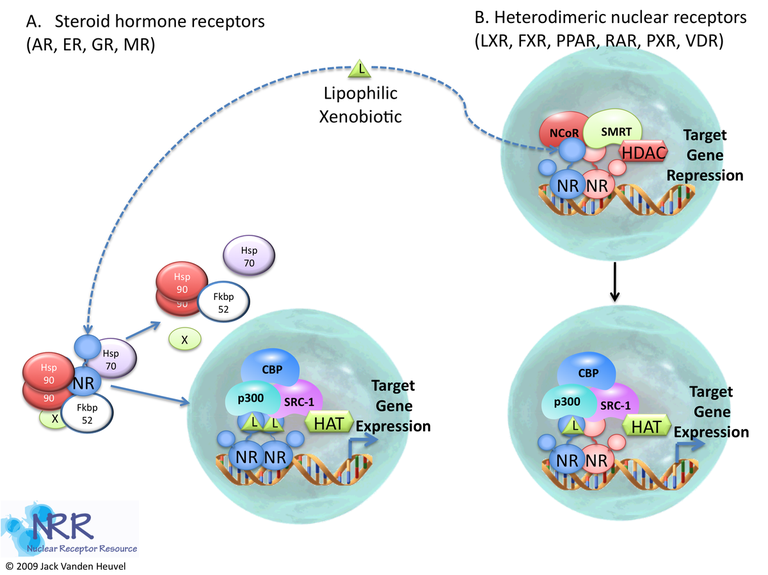
Figure 2
Prior to activation, NRs often exist in multiprotein complexes that vary depending on the family of receptor under question. When a ligand binds to its cognate receptor, a conformational change occurs ("activation") that changes the protein- protein interfaces of the molecule. As a result, the activated receptor interacts with a NR response element (NRE) within the regulatory region of a target gene; upon recruitment of various transcriptional coactivators and subsequently RNA polymerase II (polII), initiation of transcription of the target gene occurs.
Our laboratory focused on the four likely candidates for NRs that respond to dietary fatty acids, PPAR, RXR, LXR and FXR. In particular the peroxisome proliferator-activated receptors (PPARs; PPARα, β/δ and γ) are of particular interest as they serve key roles in toxicology, carcinogenesis and nutrition. We study the ligands of these receptors, the genes they affect and the physiological consequences of their activation.
Contact Us

- Professor of Molecular Toxicology
- Email jpv2@psu.edu
- Office 814-863-8532
Contact Us

- Professor of Molecular Toxicology
- Email jpv2@psu.edu
- Office 814-863-8532

